CiliaCarta: a validated compendium of ciliary genes
About the CiliaCarta
Welcome to the CiliaCarta. This webpage was created to provide easy access and search capabilities to researchers interesting in using our predictions. For detailed information please contact John van Dam.
The CiliaCarta is based on a naive Bayesian integration of a diverse set of data sets to predict novel ciliary genes. Each data set has been evaluated for its propensity to retrieve known ciliary genes and than integrated using Bayes' theorem. The CiliaCarta Score is a log transformed odds which means that a positive score indicates that a gene is more likely to be ciliary than not ciliary.
On this webpage we provide two tables:
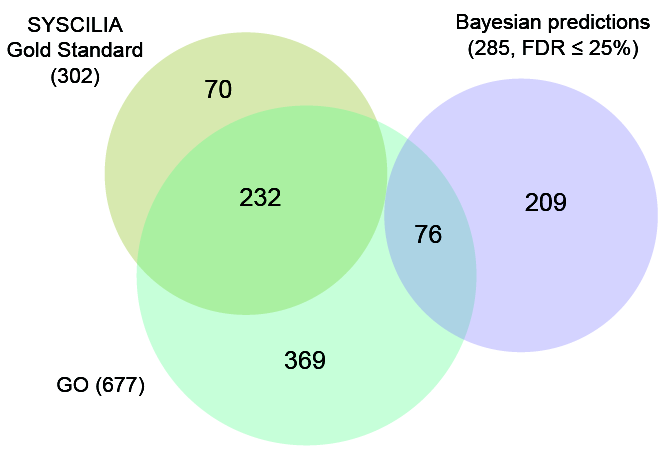
- The CiliaCarta as the compendium of ciliary genes, comprising our previous SYSCILIA Gold Standard resource, Gene Ontology annotated genes and our top predictions which have been validated experimentally.
- The naive Bayesian integration, including all datasets used in the analysis. A proteome wide list providing probabilities for each individual gene.
The selected data can directly be downloaded as a plain text *.csv file or as an excel *.xlsx file.
Citation
If you are using the CiliaCarta in your research please consider citing our paper:
We are in the process of peer review of our manuscript. A version of our manuscript can be found on the pre-print server bioRxiv:
van Dam, T. J. P. et al. CiliaCarta: An Integrated And Validated Compendium Of Ciliary Genes. PLoS One. 2019 May 16;14(5):e0216705. doi:10.1371/journal.pone.0216705
Note that we have updated this website (March 21 2018) to include recent Gene Ontology annotations. You can find the old version here.
Contact
Dr. TJP (John) van Dam
Email: t.j.p.vandam-at-uu.nl
Theoretical Biology and Bioinformatics
Utrecht University
Padualaan 8
3584 CH, Utrecht
The Netherlands
Prof. Dr. Martijn A Huynen
Email: Martijn.Huijnen-at-radboudumc.nl
CMBI, RIMLS, Radboud university medical centre
Geert Grooteplein 26-28, route 260
PO Box 9101
6500 HB Nijmegen
The Netherlands









| Ensembl Gene ID | Associated Gene Name | Description | CiliaCarta Rank | CiliaCarta Score | Inclusion criterion |
|---|---|---|---|---|---|
| Ensembl Gene ID | Associated Gene Name | Description | CiliaCarta Rank | CiliaCarta Score | Inclusion criterion |
The column 'group' denotes if a gene belonged to the positive training set (P), the negative training set (N), or unknown (G for ‘grey’). Thus in our set with an cFDR value of 25%, the 'grey' genes are our predictions for novel ciliary genes.
| Ensembl Gene ID | Associated Gene Name | Description | Group | Gene Ontology | CiliaCarta Rank | CiliaCarta Score | cFDR | TAP PPI | SILAC PPI | Mass-Spec based PPI | Y2H PPI | RFX/FOXJ1 TFBS | Cilia co-occurrence | Expression screen | Liu_2007 (CilDB) | Ross_2007 (CilDB) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ensembl Gene ID | Associated Gene Name | Description | Group | Gene Ontology | CiliaCarta Rank | CiliaCarta Score | cFDR | TAP PPI | SILAC PPI | Mass-Spec based PPI | Y2H PPI | RFX/FOXJ1 TFBS | Cilia co-occurrence | Expression screen | Liu_2007 (CilDB) | Ross_2007 (CilDB) |
This webpage uses javascripts and css from JQuery, Bootstrap and DataTables. It requires javascript to be enabled to work properly.
Funding


 The European Community's Seventh Framework Programme FP7/2009, grant agreement number 241955, SYSCILIA
The European Community's Seventh Framework Programme FP7/2009, grant agreement number 241955, SYSCILIA
 The Virgo consortium, funded by the Dutch government (FES0908), and by the Netherlands Genomics Initiative (050-060-452).
The Virgo consortium, funded by the Dutch government (FES0908), and by the Netherlands Genomics Initiative (050-060-452).